Chapter 31 System preparation for package development
Although we’ll build a very simple package, we’ll use the most modern and powerful tools for R package development. In theory, this could eventually involve compiling C/C++ code, which means you need what’s called a build environment.
31.1 Update R and RStudio
Embarking on your career as an R package developer is an important milestone. Why not celebrate by updating R and RStudio? This is something we recommended early and we recommend doing it often. Go back to Chapter 1 for reminders on the process. DO IT NOW. We are not very interested in solving problems that stem from running outdated versions of R and RStudio.
2016-11 FYI: Jenny is running R version 3.3.1 (2016-06-21) Bug in Your Hair and RStudio 1.0.44 at the time of writing.
31.2 Install devtools from CRAN
We use the devtools package to help us develop our R package. Do this:
31.3 Windows: system prep
You will probably get an immediate warning from devtools, complaining that you need Rtools in order to build R packages.
You can ignore this and successfully develop an R package that consists solely of R code. Such as our toy package.
However, we recommend you install Rtools, so you can take full advantage of devtools. Soon, you will want to use devtools::install_github() to install R packages from GitHub, instead of CRAN. You will inevitably need to build a package that includes C/C++ code, which will require Rtools.
Rtools is NOT an R package but is rather “a collection of resources for building packages for R under Microsoft Windows, or for building R itself”.
Go here and do what it says:
https://cran.r-project.org/bin/windows/Rtools/
During the installation of Rtools you will get to a window asking you to “Select Additional Tasks”. It is important that you make sure to select the box for “Edit the system PATH”.
Are we going to recommend making sure Git Bash is NOT on PATH? See #230.
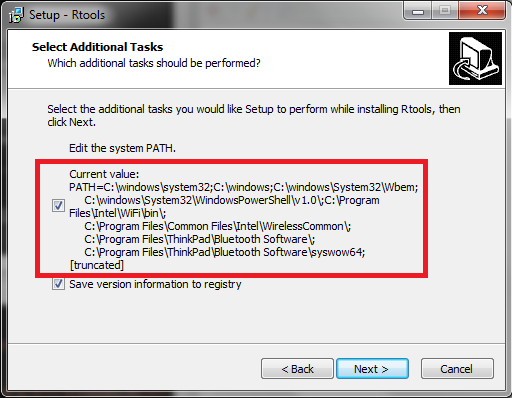
Figure 24.1: Rtools installation
After installing Rtools, restart RStudio, then do:
Hopefully you will simply see a message saying TRUE, indicating that Rtools is properly installed. But if there was a problem, you will see a longer message with next steps.
31.4 macOS: system prep
You will not get an immediate warning from devtools that you need to install anything. But before you can build R package with compiled code, you will also need to install more software. Pick one of the following:
- Minimalist approach (what I do): Install Xcode Command Line Tools.
- In the shell:
xcode-select --install
- In the shell:
- Install the current release of full Xcode from the Mac App Store. WAY more stuff than you need but advantage is App Store convenience.
- Get older or beta releases of Xcode from https://developer.apple.com/support/xcode/.
31.5 Linux: system prep
We’ve never had this section but RStudio’s devtools guide and R Packages both say the r-devel or r-base-dev package is required. What gives?
31.6 Check system prep
devtools offers a diagnostic function to check if your system is ready.
Hopefully you see TRUE!
31.7 R packages to help you build yet more R packages
Install more packages. If you already have them, update them.
- knitr
- roxygen2
- testthat
2016-11 FYI: Jenny is running these versions of these packages at the time of writing.
#> package * version date source
#> 1 devtools * 1.12.0.9000 2016-11-23 local
#> 2 knitr * 1.14.2 2016-09-07 Github (yihui/knitr@f02600d)
#> 3 roxygen2 * 5.0.1.9000 2016-10-23 Github (klutometis/roxygen@9ffbad0)
#> 4 testthat * 1.0.2.9000 2016-09-09 Github (hadley/testthat@46d15da)How to check which version of a specific package you’ve got installed:
How to install a package and all it’s dependencies:
See how profound your problem with out-of-date packages is:
Update one package:
Just update everything:
CAVEAT: The above examples will only consult your default library and default CRAN mirror. If you want to target a non-default library, use function arguments to say so. Packages that you have installed from GitHub? You’ll need to check the current-ness of your version and perform upgrades yourself.
31.8 Optional: install devtools from GitHub
We aren’t using bleeding edge features of devtools, but you could upgrade to the development version of devtools at this point.
macOS and Linux users have it easy. Do this:
For Windows instructions, see the devtools README.