Chapter 39 Use API-wrapping packages
39.1 Introduction
All this and more is described at the rOpenSci repository of R tools for interacting with the internet.
There are many ways to obtain data from the internet; let’s consider four categories:
- Click-and-download - on the internet as a “flat” file, such as CSV, XLS.
- Install-and-play - an API for which someone has written a handy R package.
- API-query - published with an unwrapped API.
- Scraping - implicit in an HTML website.
39.2 Click-and-Download
In the simplest case, the data you need is already on the internet in a tabular format. There are a couple of strategies here:
- Use
read.csvorreadr::read_csvto read the data straight into R. - Use the command line program
curlto do that work, and place it in aMakefileor shell script (see the section onmakefor more on this).
The second case is most useful when the data you want has been provided in a format that needs cleanup. For example, the World Value Survey makes several datasets available as Excel sheets. The safest option here is to download the .xls file, then read it into R with readxl::read_excel() or something similar. An exception to this is data provided as Google Spreadsheets, which can be read straight into R using the googlesheets package.
39.2.1 From rOpenSci web services page:
From rOpenSci’s CRAN Task View: Web Technologies and Services:
downloader::download()for SSL.curl::curl()for SSL.httr::GETdata read this way needs to be parsed later withread.table().rio::import()can “read a number of common data formats directly from anhttps://URL”. Isn’t that very similar to the previous?
What about packages that install data?
39.3 Data supplied on the web
Many times, the data that you want is not already organized into one or a few tables that you can read directly into R. More frequently, you find this data is given in the form of an API. Application Programming Interfaces (APIs) are descriptions of the kind of requests that can be made of a certain piece of software, and descriptions of the kind of answers that are returned.
Many sources of data – databases, websites, services – have made all (or part) of their data available via APIs over the internet. Computer programs (“clients”) can make requests of the server, and the server will respond by sending data (or an error message). This client can be many kinds of other programs or websites, including R running from your laptop.
39.4 Install-and-play
Many common web services and APIs have been “wrapped”, i.e. R functions have been written around them which send your query to the server and format the response.
Why would we want this?
- Provenance
- Reproducible
- Updating
- Ease
- Scaling
39.4.1 Load the tidyverse
39.4.2 Sightings of birds: rebird
rebird is an R interface for the eBird database. eBird lets birders upload sightings of birds, and allows everyone access to those data. rebird is on CRAN.
39.4.2.1 Search birds by geography
The eBird website categorizes some popular locations as “Hotspots”. These are areas where there are both lots of birds and lots of birders. One such location is at Iona Island, near Vancouver. You can see data for this Hotspot at http://ebird.org/ebird/hotspot/L261851.
At that link, you will see a page like this:
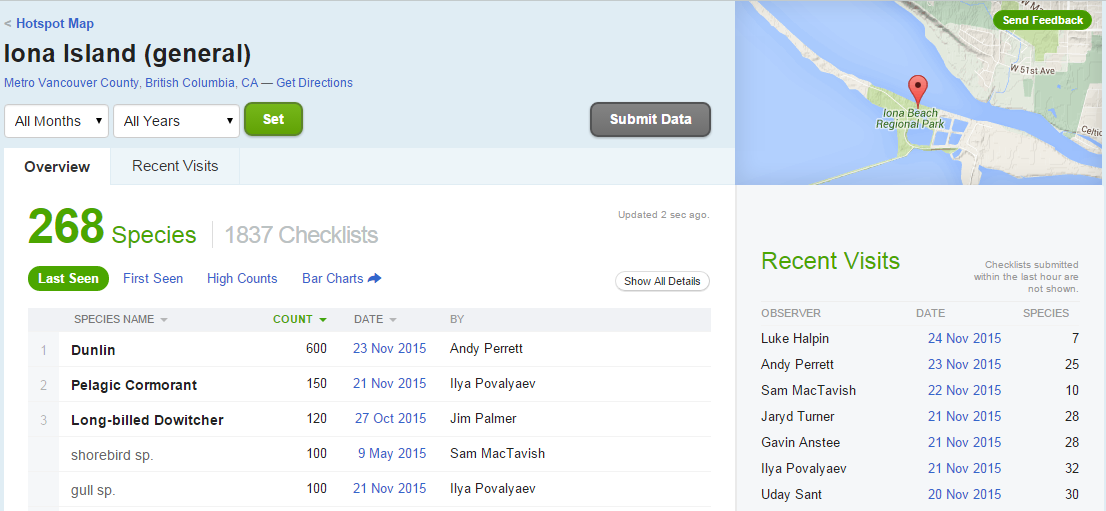
Figure 39.1: Iona Island
The data already looks to be organized in a data frame! rebird allows us to read these data directly into R (the ID code for Iona Island is “L261851”).
We can use the function ebirdgeo() to get a list for an area (note that South and West are negative):
Note: Check the defaults on this function (e.g. radius of circle, time of year).
We can also search by “region”, which refers to short codes which serve as common shorthands for different political units. For example, France is represented by the letters FR.
Find out when a bird has been seen in a certain place! Choosing a name from vanbirds above (the Bald Eagle):
eagle <- ebirdgeo(species = 'Haliaeetus leucocephalus', lat = 42, lng = -76)
eagle %>%
head() %>%
kable()rebird knows where you are:
39.4.3 Searching geographic info: geonames
rOpenSci has a package called geonames for accessing the GeoNames API. First, install the geonames package from CRAN and load it.
The geonames package website tells us that there are a few things we need to do before we can use geonames to access the GeoNames API:
- Go to the GeoNames site and create a new user account.
- Check your email and follow the instructions to activate your account.
- Click [here] to enable the free web services for your account (Note! You must be logged into your GeoNames account already for the link to work).
- Tell R your GeoNames username.
To do the last step, we could run this line in R…
…but this is insecure. We don’t want to risk committing this line and pushing it to our public GitHub page!
Instead, we can add this line to our .Rprofile so it will be hidden. One way to edit your .Rprofile is with the helper function edit_r_profile() from the usethis package. Install/load the usethis package and run edit_r_profile() in the R Console:
This will open up your .Rprofile file. Add options(geonamesUsername="my_user_name") on a new line (replace “my_user_name” with your GeoNames username).
Important: Make sure your .Rprofile ends with a blank line!
Save the file, close it, and restart R. Now we’re ready to start using geonames to search the GeoNames API.
(Also see the Cache credentials for HTTPS chapter of Happy Git and GitHub for the useR.)
39.4.3.1 Using GeoNames
What can we do? We can get access to lots of geographical information via the various GeoNames WebServices.
This countryInfo dataset is very helpful for accessing the rest of the data because it gives us the standardized codes for country and language.
39.4.3.2 Remixing geonames and rebird:
What are the cities of France?
39.4.4 Wikipedia searching
We can use geonames to search for georeferenced Wikipedia articles. Here are those within 20 km of Rio de Janerio, comparing results for English-language Wikipedia (lang = "en") and Portuguese-language Wikipedia (lang = "pt"):
39.4.5 Searching the Public Library of Science: rplos
PLOS ONE is an open-access journal. They allow access to an impressive range of search tools, and allow you to obtain the full text of their articles. rOpenSci has a package called rplos that we can use to interact with the PLOS API. They have a nice tutorial on the rOpenSci website that you can see here. First, install/load the rplos package from CRAN.
39.4.5.1 Searching PLOS ONE
Let’s follow along with the rOpenSci tutorial and do some searches:
searchplos(q= "Helianthus", fl= "id", limit = 5)
#> $meta
#> # A tibble: 1 x 2
#> numFound start
#> <int> <int>
#> 1 588 0
#>
#> $data
#> # A tibble: 5 x 1
#> id
#> <chr>
#> 1 10.1371/journal.pone.0198869
#> 2 10.1371/journal.pone.0213065
#> 3 10.1371/journal.pone.0148280
#> 4 10.1371/journal.pone.0111982
#> 5 10.1371/journal.pone.0139188searchplos("materials_and_methods:France", fl = "title, materials_and_methods")
#> $meta
#> # A tibble: 1 x 2
#> numFound start
#> <int> <int>
#> 1 15070 0
#>
#> $data
#> # A tibble: 10 x 2
#> title materials_and_methods
#> <chr> <chr>
#> 1 Population Structure in Naegleri… "\nSampling\nThe 47 N. fowleri strain…
#> 2 Borna Disease Virus Phosphoprote… "\nEthics statement\nHuman fetuses we…
#> 3 Prevalent hepatitis B surface an… "\nStudy site\nThe study was carried …
#> 4 Immunomodulation Stimulates the … "\nAnimals and tissues\nAll procedure…
#> 5 Human Neural Cells Transiently E… "\nBiological samples\nTwelve human e…
#> 6 Immunity Traits in Pigs: Substan… "\n Ethics Statement\n …
#> 7 The Plasminogen Activation Syste… "\n Differentiation of ESCs\n …
#> 8 Mesenchymal Stem Cells Repress T… "\n Ethics Statement\n …
#> 9 Adhesion of Neisseria meningitid… "\n Ethics statement\n …
#> 10 Neonatal Hyperglycemia Inhibits … "\nEthics\nAll experimental protocols…searchplos("materials_and_methods:study site", fl = "title, materials_and_methods")
#> $meta
#> # A tibble: 1 x 2
#> numFound start
#> <int> <int>
#> 1 103774 0
#>
#> $data
#> # A tibble: 10 x 2
#> title materials_and_methods
#> <chr> <chr>
#> 1 Whi5 Regulation by Site Specific… "\nYeast Culture and Strains\nCells w…
#> 2 Remote Source Document Verificat… "Two NIH-sponsored clinical trial net…
#> 3 Variance Component Analysis of a… "\n1) The study\nAddona et al. [6] co…
#> 4 Obtaining Valid Laboratory Data … "The study was a randomised, double-b…
#> 5 Transposable Prophage Mu Is Orga… "\nStrain construction\nAll strains u…
#> 6 Retention in Care and Outpatient… "\nEthics Statement\nThe Boston Unive…
#> 7 Structural and functional dissec… "\nBacterial strains and culture cond…
#> 8 Spatiotemporal Effects of Supple… "\n(a) Ethics Statement\nNo animals w…
#> 9 A-Site mRNA Cleavage Is Not Requ… "\nBacterial strains and plasmids\nAl…
#> 10 Global migration of clinical res… "\nPrimary data source\nOur primary d…searchplos("*:*", fl = "id")
#> $meta
#> # A tibble: 1 x 2
#> numFound start
#> <int> <int>
#> 1 2262279 0
#>
#> $data
#> # A tibble: 10 x 1
#> id
#> <chr>
#> 1 10.1371/journal.pone.0058099/materials_and_methods
#> 2 10.1371/journal.pone.0030394/introduction
#> 3 10.1371/journal.pone.0030394/results_and_discussion
#> 4 10.1371/journal.pone.0002157/materials_and_methods
#> 5 10.1371/journal.pone.0030394/supporting_information
#> 6 10.1371/journal.pone.0044137/materials_and_methods
#> 7 10.1371/journal.pone.0113465/materials_and_methods
#> 8 10.1371/journal.pone.0099112/introduction
#> 9 10.1371/journal.pone.0099112/results_and_discussion
#> 10 10.1371/journal.pone.0099112/materials_and_methodsHere is a list of options for the search or you can run data(plosfields) followed by plosfields in the R Console.
39.4.5.2 Take a highbrow look!
The highplos() function does “highlighted searches on PLOS Journals full-text content”.
We can then pass this output to highbrow(), which will open up our default browser where we can browse the highlighted fragments. When we run highbrow(highlighted) in our R Console this is what we see in our browser:

Figure 39.2: Example rplos highlights
39.4.5.3 Plots over time
We can use the plot_throughtime() function to visualize the results of a search over time.

39.4.6 Is it a boy or a girl? gender-associated names throughout US history
The gender package allows you access to data on the gender of names in the US. Because names change gender over the years, the probability of a name belonging to a man or a woman also depends on the year.
First, install/load the gender package from CRAN. You may be prompted to also install the companion package, genderdata. Go ahead and say yes. If you don’t see this message no need to worry, it is a one-time install.
Let’s do some searches for the name Kelsey.
gender("Kelsey")
#> # A tibble: 1 x 6
#> name proportion_male proportion_female gender year_min year_max
#> <chr> <dbl> <dbl> <chr> <dbl> <dbl>
#> 1 Kelsey 0.0314 0.969 female 1932 2012
gender("Kelsey", years = 1940)
#> # A tibble: 1 x 6
#> name proportion_male proportion_female gender year_min year_max
#> <chr> <dbl> <dbl> <chr> <dbl> <dbl>
#> 1 Kelsey 1 0 male 1940 1940